Пример машинного обучения
Ирэн Риверо (30 декабря 2022 г.)
Практический проект Прогнозирование риска рака шейки матки с помощью машинного обучения разделен на следующие задачи:
- Понять постановку проблемы и бизнес-кейс
- Импорт библиотек/наборов данных
- Выполните исследовательский анализ данных
- Выполнение визуализации данных
- Подготовьте данные перед обучением модели
- Обучение и оценка модели XG-Boost
Спасибо профессору Райану Ахмеду за то, что он сделал этот проект таким простым!
XGBoost
XGBoost или Extreme Gradient Boosting — это алгоритм, который выбирают многие специалисты по данным, и его можно использовать для задач регрессии и классификации. XGBoost — это алгоритм обучения под наблюдением, реализующий алгоритм деревьев с градиентным усилением.
Алгоритм работает путем объединения ансамбля прогнозов из нескольких слабых моделей. Он устойчив ко многим распределениям данных и отношениям и предлагает множество гиперпараметров для настройки производительности модели. XGBoost предлагает повышенную скорость и улучшенное использование памяти.
Усиление работает за счет обучения на предыдущих ошибках (ошибки в прогнозах модели) для создания лучших прогнозов на будущее. Повышение — это метод ансамблевого машинного обучения, который работает путем последовательного обучения слабых моделей.
Каждая модель пытается извлечь уроки из предыдущей слабой модели и стать лучше в прогнозировании. Алгоритмы повышения работают, строя модель из обучающих данных, затем вторая модель строится на основе ошибок (остатков) первой модели. Алгоритм повторяется до тех пор, пока не будет создано максимальное количество моделей или пока модель не будет давать хорошие прогнозы.
Понять формулировку проблемы
В этом практическом проекте мы создадим и обучим модель XGBoost для прогнозирования рака шейки маткиу 858 пациентов.
Набор данных был собран в «Университетской больнице Каракаса» в Каракасе, Венесуэла, и содержит демографическую информацию, привычки и исторические медицинские записи 858 пациентов.
Рак шейки матки убивает около 4 000 женщин в США и около 300 000 женщин во всем мире. Благодаря усилению медицинского скрининга смертность от рака шейки матки с 1955 по 1992 год снизилась на 74%.
Исследования показали, что высокая сексуальная активность и вирус папилломы человека (ВПЧ) являются одним из ключевых факторов, повышающих риск развития рака шейки матки.
Наличие гормонов в оральных контрацептивах, многодетность и курение повышают риск развития рака шейки матки, особенно у женщин, инфицированных ВПЧ. Кроме того, люди со слабой иммунной системой (ВИЧ/СПИД) имеют высокий риск заражения ВПЧ.

Импорт набора данных и библиотек
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import zipfile
!pip install jupyterthemes
!pip install plotly
import plotly.express as px
from jupyterthemes import jtplot
jtplot.style(theme = 'monokai', context = 'notebook', ticks = True, grid = False)
# setting the style of the notebook to be monokai theme
# this line of code is important to ensure that we are able to see the x and y axes clearly
# If you don't run this code line, you will notice that the xlabel and ylabel on any plot is black on black and it will be hard to see them.
# import the csv files using pandas
cancer_df = pd.read_csv('cervical_cancer.csv')
# (int) Age
# (int) Number of sexual partners
# (int) First sexual intercourse (age)
# (int) Num of pregnancies
# (bool) Smokes
# (bool) Smokes (years)
# (bool) Smokes (packs/year)
# (bool) Hormonal Contraceptives
# (int) Hormonal Contraceptives (years)
# (bool) IUD ("IUD" stands for "intrauterine device" and used for birth control
# (int) IUD (years)
# (bool) STDs (Sexually transmitted disease)
# (int) STDs (number)
# (bool) STDs:condylomatosis
# (bool) STDs:cervical condylomatosis
# (bool) STDs:vaginal condylomatosis
# (bool) STDs:vulvo-perineal condylomatosis
# (bool) STDs:syphilis
# (bool) STDs:pelvic inflammatory disease
# (bool) STDs:genital herpes
# (bool) STDs:molluscum contagiosum
# (bool) STDs:AIDS
# (bool) STDs:HIV
# (bool) STDs:Hepatitis B
# (bool) STDs:HPV
# (int) STDs: Number of diagnosis
# (int) STDs: Time since first diagnosis
# (int) STDs: Time since last diagnosis
# (bool) Dx:Cancer
# (bool) Dx:CIN
# (bool) Dx:HPV
# (bool) Dx
# (bool) Hinselmann: target variable - A colposcopy is a procedure in which doctors examine the cervix.
# (bool) Schiller: target variable - Schiller's Iodine test is used for cervical cancer diagnosis
# (bool) Cytology: target variable - Cytology is the exam of a single cell type used for cancer screening.
# (bool) Biopsy: target variable - Biopsy is performed by removing a piece of tissue and examine it under microscope,
# Biopsy is the main way doctors diagnose most types of cancer.
# Let's explore the dataframe
cancer_df

#Print the last 20 rows in the dataframe cancer_df.tail(20) #Print the first 20 rows in the dataframe cancer_df.head(20)

Выполнение исследовательского анализа данных
# Get data frame info cancer_df.info() <class 'pandas.core.frame.DataFrame'> RangeIndex: 858 entries, 0 to 857 Data columns (total 36 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 Age 858 non-null int64 1 Number of sexual partners 858 non-null object 2 First sexual intercourse 858 non-null object 3 Num of pregnancies 858 non-null object 4 Smokes 858 non-null object 5 Smokes (years) 858 non-null object 6 Smokes (packs/year) 858 non-null object 7 Hormonal Contraceptives 858 non-null object 8 Hormonal Contraceptives (years) 858 non-null object 9 IUD 858 non-null object 10 IUD (years) 858 non-null object 11 STDs 858 non-null object 12 STDs (number) 858 non-null object 13 STDs:condylomatosis 858 non-null object 14 STDs:cervical condylomatosis 858 non-null object 15 STDs:vaginal condylomatosis 858 non-null object 16 STDs:vulvo-perineal condylomatosis 858 non-null object 17 STDs:syphilis 858 non-null object 18 STDs:pelvic inflammatory disease 858 non-null object 19 STDs:genital herpes 858 non-null object 20 STDs:molluscum contagiosum 858 non-null object 21 STDs:AIDS 858 non-null object 22 STDs:HIV 858 non-null object 23 STDs:Hepatitis B 858 non-null object 24 STDs:HPV 858 non-null object 25 STDs: Number of diagnosis 858 non-null int64 26 STDs: Time since first diagnosis 858 non-null object 27 STDs: Time since last diagnosis 858 non-null object 28 Dx:Cancer 858 non-null int64 29 Dx:CIN 858 non-null int64 30 Dx:HPV 858 non-null int64 31 Dx 858 non-null int64 32 Hinselmann 858 non-null int64 33 Schiller 858 non-null int64 34 Citology 858 non-null int64 35 Biopsy 858 non-null int64 dtypes: int64(10), object(26) memory usage: 241.4+ KB # Get the statistics of the data frame cancer_df.describe()

# Notice many question marks indicating missing values cancer_df

# Let's replace '?' with NaN
cancer_df = cancer_df.replace('?', np.nan)
cancer_df
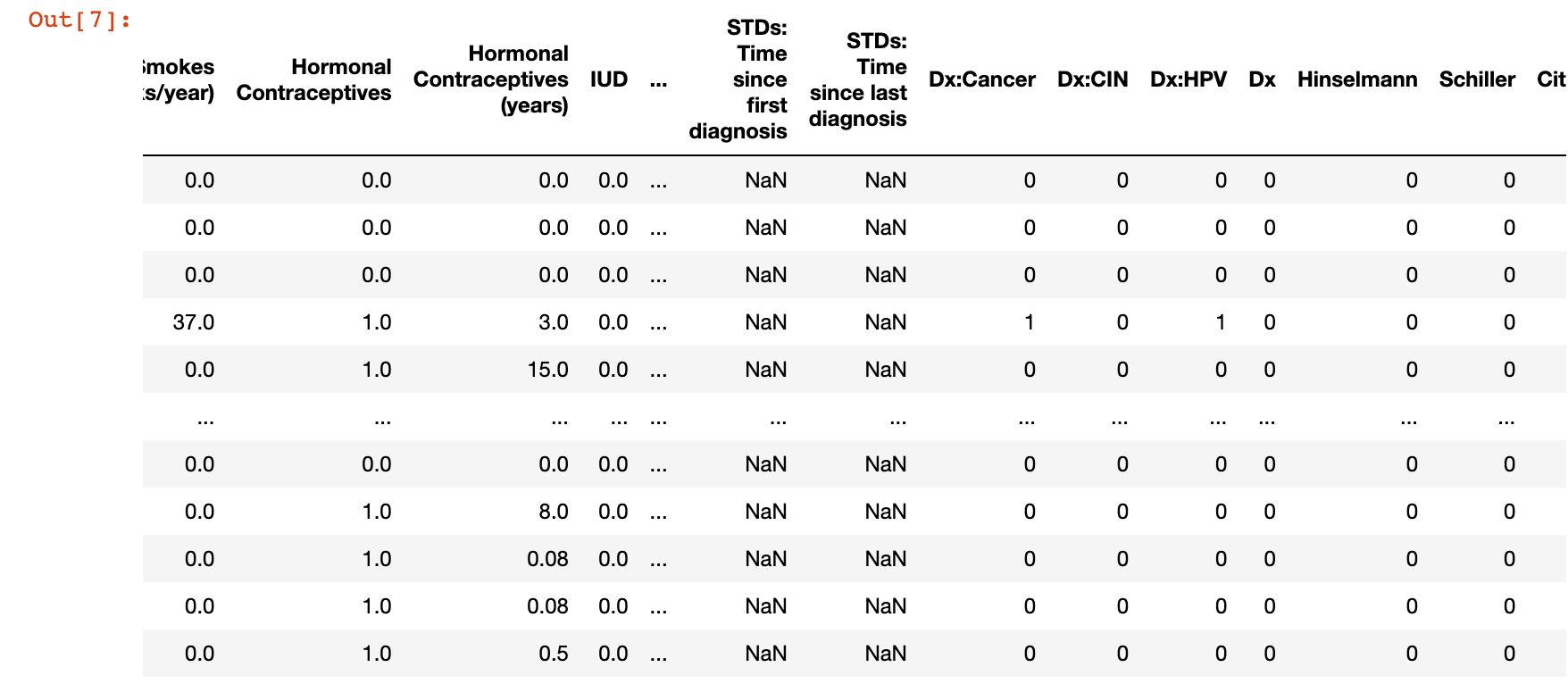
cancer_df.isnull()

# Plot heatmap plt.figure(figsize = (20, 20)) sns.heatmap(cancer_df.isnull())

# Get data frame info cancer_df.info() <class 'pandas.core.frame.DataFrame'> RangeIndex: 858 entries, 0 to 857 Data columns (total 36 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 Age 858 non-null int64 1 Number of sexual partners 832 non-null object 2 First sexual intercourse 851 non-null object 3 Num of pregnancies 802 non-null object 4 Smokes 845 non-null object 5 Smokes (years) 845 non-null object 6 Smokes (packs/year) 845 non-null object 7 Hormonal Contraceptives 750 non-null object 8 Hormonal Contraceptives (years) 750 non-null object 9 IUD 741 non-null object 10 IUD (years) 741 non-null object 11 STDs 753 non-null object 12 STDs (number) 753 non-null object 13 STDs:condylomatosis 753 non-null object 14 STDs:cervical condylomatosis 753 non-null object 15 STDs:vaginal condylomatosis 753 non-null object 16 STDs:vulvo-perineal condylomatosis 753 non-null object 17 STDs:syphilis 753 non-null object 18 STDs:pelvic inflammatory disease 753 non-null object 19 STDs:genital herpes 753 non-null object 20 STDs:molluscum contagiosum 753 non-null object 21 STDs:AIDS 753 non-null object 22 STDs:HIV 753 non-null object 23 STDs:Hepatitis B 753 non-null object 24 STDs:HPV 753 non-null object 25 STDs: Number of diagnosis 858 non-null int64 26 STDs: Time since first diagnosis 71 non-null object 27 STDs: Time since last diagnosis 71 non-null object 28 Dx:Cancer 858 non-null int64 29 Dx:CIN 858 non-null int64 30 Dx:HPV 858 non-null int64 31 Dx 858 non-null int64 32 Hinselmann 858 non-null int64 33 Schiller 858 non-null int64 34 Citology 858 non-null int64 35 Biopsy 858 non-null int64 dtypes: int64(10), object(26) memory usage: 241.4+ KB # Since STDs: Time since first diagnosis and STDs: Time since last diagnosis have more than 80% missing values # we can drop them cancer_df = cancer_df.drop(columns = ['STDs: Time since first diagnosis', 'STDs: Time since last diagnosis']) cancer_df

# Since most of the column types are object, we are not able to get the statistics of the dataframe. # Convert them to numeric type cancer_df = cancer_df.apply(pd.to_numeric) cancer_df.info() <class 'pandas.core.frame.DataFrame'> RangeIndex: 858 entries, 0 to 857 Data columns (total 34 columns): # Column Non-Null Count Dtype --- ------ -------------- ----- 0 Age 858 non-null int64 1 Number of sexual partners 832 non-null float64 2 First sexual intercourse 851 non-null float64 3 Num of pregnancies 802 non-null float64 4 Smokes 845 non-null float64 5 Smokes (years) 845 non-null float64 6 Smokes (packs/year) 845 non-null float64 7 Hormonal Contraceptives 750 non-null float64 8 Hormonal Contraceptives (years) 750 non-null float64 9 IUD 741 non-null float64 10 IUD (years) 741 non-null float64 11 STDs 753 non-null float64 12 STDs (number) 753 non-null float64 13 STDs:condylomatosis 753 non-null float64 14 STDs:cervical condylomatosis 753 non-null float64 15 STDs:vaginal condylomatosis 753 non-null float64 16 STDs:vulvo-perineal condylomatosis 753 non-null float64 17 STDs:syphilis 753 non-null float64 18 STDs:pelvic inflammatory disease 753 non-null float64 19 STDs:genital herpes 753 non-null float64 20 STDs:molluscum contagiosum 753 non-null float64 21 STDs:AIDS 753 non-null float64 22 STDs:HIV 753 non-null float64 23 STDs:Hepatitis B 753 non-null float64 24 STDs:HPV 753 non-null float64 25 STDs: Number of diagnosis 858 non-null int64 26 Dx:Cancer 858 non-null int64 27 Dx:CIN 858 non-null int64 28 Dx:HPV 858 non-null int64 29 Dx 858 non-null int64 30 Hinselmann 858 non-null int64 31 Schiller 858 non-null int64 32 Citology 858 non-null int64 33 Biopsy 858 non-null int64 dtypes: float64(24), int64(10) memory usage: 228.0 KB # Get the statistics of the dataframe cancer_df.describe()

cancer_df.mean() Age 26.820513 Number of sexual partners 2.527644 First sexual intercourse 16.995300 Num of pregnancies 2.275561 Smokes 0.145562 Smokes (years) 1.219721 Smokes (packs/year) 0.453144 Hormonal Contraceptives 0.641333 Hormonal Contraceptives (years) 2.256419 IUD 0.112011 IUD (years) 0.514804 STDs 0.104914 STDs (number) 0.176627 STDs:condylomatosis 0.058433 STDs:cervical condylomatosis 0.000000 STDs:vaginal condylomatosis 0.005312 STDs:vulvo-perineal condylomatosis 0.057105 STDs:syphilis 0.023904 STDs:pelvic inflammatory disease 0.001328 STDs:genital herpes 0.001328 STDs:molluscum contagiosum 0.001328 STDs:AIDS 0.000000 STDs:HIV 0.023904 STDs:Hepatitis B 0.001328 STDs:HPV 0.002656 STDs: Number of diagnosis 0.087413 Dx:Cancer 0.020979 Dx:CIN 0.010490 Dx:HPV 0.020979 Dx 0.027972 Hinselmann 0.040793 Schiller 0.086247 Citology 0.051282 Biopsy 0.064103 dtype: float64 # Replace null values with mean cancer_df = cancer_df.fillna(cancer_df.mean()) cancer_df

# Nan heatmap plt.figure(figsize = (20,20)) sns.heatmap(cancer_df.isnull(), yticklabels = False)

#What is the age range of people involved in this study? #What are the biopsy results for the oldest person in this study? cancer_df['Age'].min() 13 cancer_df['Age'].max() 84 cancer_df[cancer_df['Age'] == 84]

Выполнить визуализацию данных
# Get the correlation matrix corr_matrix=cancer_df.corr() corr_matrix

# Plot the correlation matrix plt.figure(figsize = (30, 30)) sns.heatmap (corr_matrix, annot =True) plt.show()

fig = px.bar(cancer_df, x="Age", y="Biopsy", orientation='h', color = 'Age', labels = dict(Age = 'Total Instances')) fig.show()

#Plot the histogram for the entire DataFrame cancer_df.hist(bins=10, figsize = (30, 30), color= 'b')

Подготовьте данные перед обучением
target_df = cancer_df ['Biopsy']
input_df = cancer_df.drop(columns = ['Biopsy'])
target_df.shape
(858,)
input_df.shape
(858, 33)
target_df
0 0
1 0
2 0
3 0
4 0
..
853 0
854 0
855 0
856 0
857 0
Name: Biopsy, Length: 858, dtype: int64
X = np.array(input_df).astype('float32')
y = np.array(target_df).astype('float32')
# reshaping the array from (421570,) to (421570, 1)
# y = y.reshape(-1,1)
y.shape
(858,)
# scaling the data before feeding the model
from sklearn.preprocessing import StandardScaler, MinMaxScaler
scaler = StandardScaler()
X = scaler.fit_transform(X)
X
array([[-1.0385634e+00, 8.9706147e-01, -7.1509570e-01, ...,
-2.0622157e-01, -3.0722591e-01, -2.3249528e-01],
[-1.3917956e+00, -9.3074709e-01, -1.0734857e+00, ...,
-2.0622157e-01, -3.0722591e-01, -2.3249528e-01],
[ 8.4534228e-01, -9.3074709e-01, 2.2945171e-07, ...,
-2.0622157e-01, -3.0722591e-01, -2.3249528e-01],
...,
[-2.1435463e-01, -3.2147753e-01, 1.6845580e-03, ...,
-2.0622157e-01, -3.0722591e-01, 4.3011627e+00],
[ 7.2759819e-01, -3.2147753e-01, 2.5104153e+00, ...,
-2.0622157e-01, -3.0722591e-01, -2.3249528e-01],
[ 2.5662178e-01, -3.2147753e-01, 1.0768549e+00, ...,
-2.0622157e-01, -3.0722591e-01, -2.3249528e-01]], dtype=float32)
# scaler = StandardScaler()
# X_train = scaler.fit_transform(X_train)
# X_test = scaler.transform(X_test)
# X_val = scaler.transform(X_val)
# spliting the data in to test and train sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
X_test, X_val, y_test, y_val = train_test_split(X_test, y_test, test_size = 0.5)
#Split the data such that the testing data is quarter the size of the training data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25)
Обучение и оценка классификатора XGBoost
!pip install xgboost
# Train an XGBoost classifier model
import xgboost as xgb
model = xgb.XGBClassifier(learning_rate = 0.1, max_depth = 5, n_estimators = 10)
model.fit(X_train, y_train)
result_train = model.score(X_train, y_train)
print("Accuracy : {}".format(result_train))
Accuracy : 0.9860031104199067
# predict the score of the trained model using the testing dataset
result_test = model.score(X_test, y_test)
print("Accuracy : {}".format(result_test))
Accuracy : 0.9441860465116279
# make predictions on the test data
y_predict = model.predict(X_test)
from sklearn.metrics import confusion_matrix, classification_report
print(classification_report(y_test, y_predict))
precision recall f1-score support
0.0 0.95 0.99 0.97 194
1.0 0.85 0.52 0.65 21
accuracy 0.94 215
macro avg 0.90 0.76 0.81 215
weighted avg 0.94 0.94 0.94 215
cm = confusion_matrix(y_predict, y_test)
sns.heatmap(cm, annot = True)

plt.figure(figsize=(10, 10))
cm = confusion_matrix(y_predict, y_test)
sns.heatmap(cm, annot = True,fmt = '.2f')
plt.ylabel('Predicted class')
plt.xlabel('Actual class')

#Retrain the model with 10x and 100x the number of estimators and tree depth
#Plot the confusion matrix
#Comment on the performance of the mode
model = xgb.XGBClassifier(learning_rate = 0.1, max_depth = 50, n_estimators = 100)
model.fit(X_train, y_train)
result_train = model.score(X_train, y_train)
print("Accuracy : {}".format(result_train))
Accuracy : 0.9956268221574344
# predict the score of the trained model using the testing dataset
result = model.score(X_test, y_test)
print("Accuracy : {}".format(result))
Accuracy : 0.9651162790697675
# make predictions on the test data
y_predict = model.predict(X_test)
from sklearn.metrics import confusion_matrix, classification_report
print(classification_report(y_test, y_predict))
precision recall f1-score support
0.0 0.99 0.97 0.98 80
1.0 0.71 0.83 0.77 6
accuracy 0.97 86
macro avg 0.85 0.90 0.88 86
weighted avg 0.97 0.97 0.97 86
plt.figure(figsize=(10, 10))
cm = confusion_matrix(y_predict, y_test)
sns.heatmap(cm, annot = True,fmt = '.2f')
plt.ylabel('Predicted class')
plt.xlabel('Actual class')

Попытка получить лучшую поддержку
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25)
model = xgb.XGBClassifier(learning_rate = 0.1, max_depth = 50, n_estimators = 100)
model.fit(X_train, y_train)
XGBClassifier(max_depth=50)
result_train = model.score(X_train, y_train)
print("Accuracy : {}".format(result_train))
Accuracy : 1.0
result = model.score(X_test, y_test)
print("Accuracy : {}".format(result))
Accuracy : 0.9627906976744186
y_predict = model.predict(X_test)
from sklearn.metrics import confusion_matrix, classification_report
print(classification_report(y_test, y_predict))
precision recall f1-score support
0.0 0.98 0.98 0.98 205
1.0 0.60 0.60 0.60 10
accuracy 0.96 215
macro avg 0.79 0.79 0.79 215
weighted avg 0.96 0.96 0.96 215
plt.figure(figsize=(10, 10))
cm = confusion_matrix(y_predict, y_test)
sns.heatmap(cm, annot = True,fmt = '.2f')
plt.ylabel('Predicted class')
plt.xlabel('Actual class')
